This web page was produced as an assignment for Genetics 564, an undergraduate course at the University of Wisconsin-Madison.
Protein Domains
What are protein domains?
Domains are distinct functional and/or structural units in a protein. Each domain is usually responsible for a specific function that contributes to the overall role of the protein within a cell [1]. Obtaining information about the domains contained in a protein of interest can lead to information about the proteins function. There are many databases available that give information about the domains in a protein. For example: Pfam, SMART or Prosite.
Domains are distinct functional and/or structural units in a protein. Each domain is usually responsible for a specific function that contributes to the overall role of the protein within a cell [1]. Obtaining information about the domains contained in a protein of interest can lead to information about the proteins function. There are many databases available that give information about the domains in a protein. For example: Pfam, SMART or Prosite.
Domains in CNGA3 protein?

Amino Acids 206-399: The ion transport domain contains Sodium, Potassium, and Calcium ion channels. It has 6 transmembrane helices [1].

Amino Acids 502-593: The cNMP binding domain is a region that binds cNMP, as the name implies [2].
Domains in CNGA3 protein homologs:
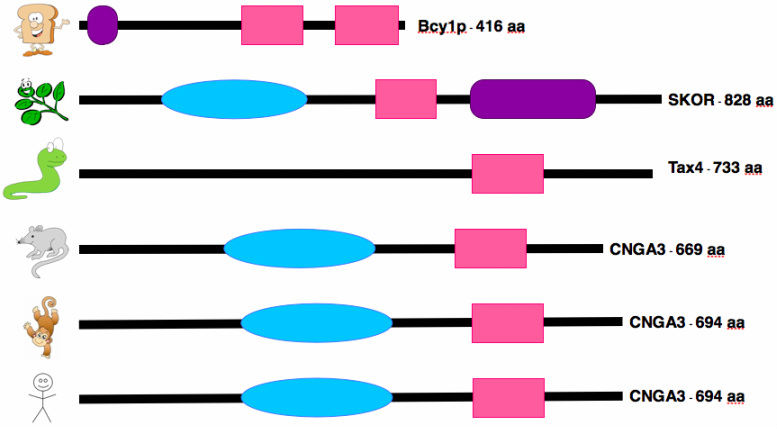
The figure below shows where the above two domains are located on the CNGA3 protein in other model organisms.
The figure below shows where the above two domains are located on the CNGA3 protein in other model organisms.
Discussion:
Both of the domains found in the human CNGA3 protein are found in similar size and location in many other model organisms with eyes, as shown (mouse, chimp). The domains are less similar in organisms without eyes (shown: yeast, arabadopsis, C. elegans), but still present in all cases except for C. elegans missing the ion transport domain. The cNMP binding domain is more highly conserved than the ion transport domain across species. This is what I expected, but I was still surprised at just how well conserved the domains were in species without eyes.
Both of the domains found in the human CNGA3 protein are found in similar size and location in many other model organisms with eyes, as shown (mouse, chimp). The domains are less similar in organisms without eyes (shown: yeast, arabadopsis, C. elegans), but still present in all cases except for C. elegans missing the ion transport domain. The cNMP binding domain is more highly conserved than the ion transport domain across species. This is what I expected, but I was still surprised at just how well conserved the domains were in species without eyes.
References:
[1] http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
[2] Pfam. Domains within Homo sapiens protein CNGA3_Human. retrieved May 16, 2014. http://pfam.xfam.org/search/sequence
[1] http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
[2] Pfam. Domains within Homo sapiens protein CNGA3_Human. retrieved May 16, 2014. http://pfam.xfam.org/search/sequence