This web page was produced as an assignment for Genetics 564, an undergraduate course at the University of Wisconsin-Madison.
Gene - Homology
What is homology?
Homology indicates a structure that is passed down through a common ancestor to various species. In regards to this page homology is the existence of common ancestry between the CNGA-3 gene in the species listed below [1]. Homology can be determined by using the genes sequence and comparing how closely related this sequence is in different species. A good starting place when looking for homologs is a website called HomoloGene. After looking at these species it is important check the accuracy with another website that utilizes a tool called BLAST. Blast is also a good tool to find additional homologous or can be used as a starting point.
BLAST is used to compare the mRNA sequence in homo sapiens (humans) to other sequences that have been entered in the database. This search yields many different species and genes that are to some extent identical to the homo sapien sequence that was originally entered [2]. The BLAST website can also provide links to the FASTA for the gene in the various species. This lays out the sequence of the gene and can later be useful in depicting the phylogeny of the species. For more information on phylogeny click here.
Homology indicates a structure that is passed down through a common ancestor to various species. In regards to this page homology is the existence of common ancestry between the CNGA-3 gene in the species listed below [1]. Homology can be determined by using the genes sequence and comparing how closely related this sequence is in different species. A good starting place when looking for homologs is a website called HomoloGene. After looking at these species it is important check the accuracy with another website that utilizes a tool called BLAST. Blast is also a good tool to find additional homologous or can be used as a starting point.
BLAST is used to compare the mRNA sequence in homo sapiens (humans) to other sequences that have been entered in the database. This search yields many different species and genes that are to some extent identical to the homo sapien sequence that was originally entered [2]. The BLAST website can also provide links to the FASTA for the gene in the various species. This lays out the sequence of the gene and can later be useful in depicting the phylogeny of the species. For more information on phylogeny click here.
Examples of homology in the CNGA3 gene:
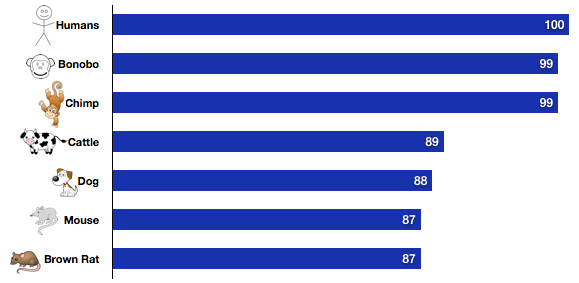
Below is a graph that shows examples of homologous in regards to the cyclic nucleotide gate alpha-3 gene. Because the CNGA-3 gene is so long the mRNA was used when searching for homologous species. The information presented below was obtained through BLAST. After determining what the homologous genes are research can be gathered on the homologs to gain more insight into the function and properties of the CNGA3 gene. It can also help guide future research questions that could helo learn more about Achromatopsia. Humans are difficult to study, so finding homology in other organisms, especially model organisms (such as the mouse), shows which species can be studied and possibly applied to benefit humans. Below the graph each specie is listed with its accession number (useful to study the protein in that species) and link to its FASTA page.
Below is a graph that shows examples of homologous in regards to the cyclic nucleotide gate alpha-3 gene. Because the CNGA-3 gene is so long the mRNA was used when searching for homologous species. The information presented below was obtained through BLAST. After determining what the homologous genes are research can be gathered on the homologs to gain more insight into the function and properties of the CNGA3 gene. It can also help guide future research questions that could helo learn more about Achromatopsia. Humans are difficult to study, so finding homology in other organisms, especially model organisms (such as the mouse), shows which species can be studied and possibly applied to benefit humans. Below the graph each specie is listed with its accession number (useful to study the protein in that species) and link to its FASTA page.
Discussion of the CNGA3 homologs:
All of the homologs listed below have a very high percent identity. This was expected because CNGA3s function is crucial for species that have eyes. There are various species that are homologous in regards to this mRNA, however far more homologs were found for the CNGA-3 protein. Species such as E. coli, yeast and Arabidopsis were not found as homologs. This makes sense because the gene is important to the cone receptors in the retina and those species do not have retinas. Because the protein has many more useful homologs, I have chosen to focus on the protein in regards to future studies and experiment ideas.
All of the homologs listed below have a very high percent identity. This was expected because CNGA3s function is crucial for species that have eyes. There are various species that are homologous in regards to this mRNA, however far more homologs were found for the CNGA-3 protein. Species such as E. coli, yeast and Arabidopsis were not found as homologs. This makes sense because the gene is important to the cone receptors in the retina and those species do not have retinas. Because the protein has many more useful homologs, I have chosen to focus on the protein in regards to future studies and experiment ideas.