This web page was produced as an assignment for Genetics 564, an undergraduate course at the University of Wisconsin-Madison.
Gene Ontology
|
What is Gene Ontology?
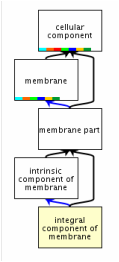
Gene ontology is a major bioinformatics project seeking to create a database that unifies representation of genes across species so that scientists can more easily interpret their experimental data [1]. This project uses a common language which allows data to be compared across species using the same vocabulary. Gene ontology uses three main categories to describe a gene: biological processes, molecular functions and cellular components. These categories are refered to as the "go" terms. The biological process terms represent collections of molecular events with a well defined beginning and end that a gene is a part of. The molecular function terms provide information regarding what jobs or abilities a gene may have. Finally, the cellular component terms tell the user where the gene of interest can be found [1]. |
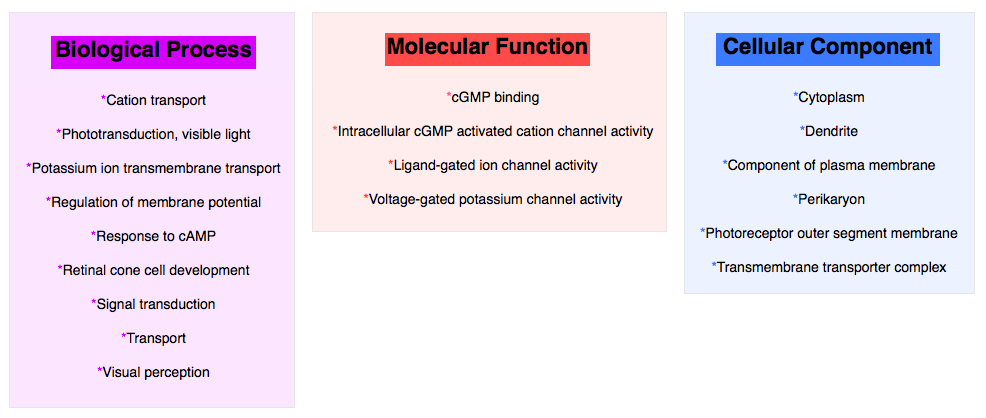
CNGA3 Ontology:
Analysis of CNGA3 Ontology:
As shown above, CNGA3 is involved in many different biological processes. These processes all reconfirm the function of CNGA3 because they are all related to vision. The information from all three categories shows the complexity of the genes function and how many smaller activities come together to ultimately allow the retina to function properly.
As shown above, CNGA3 is involved in many different biological processes. These processes all reconfirm the function of CNGA3 because they are all related to vision. The information from all three categories shows the complexity of the genes function and how many smaller activities come together to ultimately allow the retina to function properly.
References:
[1] The Gene Ontology Consortium. "Gene ontology: tool for the unification of biology." Nature Genetics. May 2000; 25(1):25-9.
[1] The Gene Ontology Consortium. "Gene ontology: tool for the unification of biology." Nature Genetics. May 2000; 25(1):25-9.